Dendrite Development
Project leader: Tabea Schilling
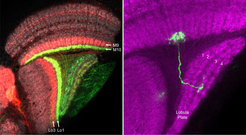
Right panel: Drosophila optic lobe with a single T4 neuron labelled with GFP (in green). The four layers of the lobula plate are visualized by staining Bruchpilot (in magenta). The axon terminal of the labelled T4 neuron is restricted to lobula plate layer 3.
How neurons acquire a diverse repertoire of structural and functional properties in order to establish working neural circuits remains poorly understood. To address this question, we use the T4/T5 neurons of the Drosophila visual system as a model because of their genetic accessibility and the extensive knowledge we have about their morphology and physiology. The dendrites of T4 and T5 neurons represent the first processing stage in the fly visual system in which the direction of image motion is encoded. While T4 neurons respond to motion of brightness increments, T5 neurons detect motion of brightness decrements (Maisak et al., 2013). This specialization results from their dendrites arborizing in different neuropil regions (T4 dendrites in the medulla, T5 dendrites in the lobula) and therefore receiving synaptic input from different neuronal types. A common attribute of T4 and T5 neurons is the layer-specific innervation displayed by their dendrites and axons (Figure 1) (Fischbach and Dittrich, 1989), which is thought to support precise synaptic connectivity. We have found that SoxN and Sox102F, two members of the SOX family of transcription factors, are required in all T4 and T5 neurons during their maturation in order to restrict their dendrites and axons into single synaptic layers. When this process fails, some postsynaptic partners of T4/T5 neurons also exhibit aberrant morphologies, which affects the proper functioning of the fly motion sensing circuits (Figure 2) (Schilling et al., 2019).
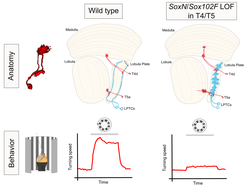
Both T4 and T5 neurons can be further subdivided into four subtypes (a,b,c,d), each innervating only one of the four lobula plate layers (Figure 1) (Fischbach and Dittrich, 1989). Furthermore, each T4/T5 subtype responds exclusively to motion in one of the four cardinal directions (Maisak et al., 2013). Differences between T4/T5 neuron subtypes in directional tuning are thought to result from differences in the spatial organization of synapses they receive from an identical set of input neurons (Serbe et al., 2016; Arenz et al., 2017; Takemura et al., 2017; Shinomiya et al., 2019). How do these differences emerge during development? Interestingly, neurons of each T4/T5 subtype orient their dendrites in a subtype-specific manner (Takemura et al., 2017; Shinomiya et al., 2019). Therefore, dendrite orientation appears strictly linked to directional tuning. Our working hypothesis is that the direction in which a T4/T5 dendrite grows during development determines the arrangement of its synaptic inputs, and thus its direction selectivity. We are currently attempting to understand the cellular rules and underlying molecular mechanisms by which the four T4/T5 neuron subtypes acquire their distinctly oriented dendritic arbors.
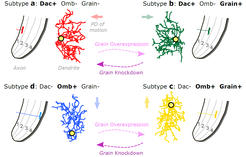
In a recent study, we investigated when and how the dendrites of the four T4/T5 subtypes acquire their specific orientations, and characterized the transcriptomes of all T4/T5 neurons during this process (Hoermann et al., 2020). We identified a simple and stable code of transcription factors defining the four T4/T5 subtypes during dendrite development. To test whether different sets of transcription factors specify distinct dendrite orientations, we manipulated them in specific T4/T5 subtypes. We found that Grain acts together with different transcription factors in subtypes b and c to coordinate dendrite and axon morphogenesis in order to differentiate their morphologies from those of subtypes a and d, respectively (Figure 3). Our study also revealed many cell-membrane proteins with subtype-specific expression patterns during dendrite growth, whose roles in dendrite morphogenesis are being investigated by loss and gain of function approaches. Elucidating how the four T4/T5 subtypes acquire their specific dendrite morphologies will further help to understand better (1) the strategies that differentiate neuron subtypes across different neuronal populations and species, and (2) the developmental programs generating structural specializations that underlie functional specializations. Furthermore, we envision that this work will provide us with tools for manipulating specific neuronal structural properties to reveal their role in neuronal computation.
References
Arenz A, Drews MS, Richter FG, Ammer G, Borst A (2017) The Temporal Tuning of the Drosophila Motion Detectors Is Determined by the Dynamics of Their Input Elements. Current biology : CB 27:929-944. DOI
Fischbach K-F, Dittrich APM (1989) The optic lobe of Drosophila melanogaster. I. A Golgi analysis of wild-type structure. Cell and Tissue Research 258:441-475.
Hoermann N, Schilling T, Haji Ali A, Serbe E, Mayer C, Borst A, Pujol Martí J (2020) A combinatorial code of transcription factors specifies subtypes of visual motion-sensing neurons in Drosophila. Development, online 3. April 2020. DOI.
Maisak MS, Haag J, Ammer G, Serbe E, Meier M, Leonhardt A, Schilling T, Bahl A, Rubin GM, Nern A, Dickson BJ, Reiff DF, Hopp E, Borst A (2013) A directional tuning map of Drosophila elementary motion detectors. Nature 500:212-216. DOI
Schilling T, Ali AH, Leonhardt A, Borst A, Pujol-Marti J (2019) Transcriptional control of morphological properties of direction-selective T4/T5 neurons in Drosophila. Development (Cambridge, England) 146. DOI
Serbe E, Meier M, Leonhardt A, Borst A (2016) Comprehensive Characterization of the Major Presynaptic Elements to the Drosophila OFF Motion Detector. Neuron 89:829-841. DOI
Shinomiya K et al. (2019) Comparisons between the ON- and OFF-edge motion pathways in the Drosophila brain. eLife 8. DOI
Takemura SY, Nern A, Chklovskii DB, Scheffer LK, Rubin GM, Meinertzhagen IA (2017) The comprehensive connectome of a neural substrate for 'ON' motion detection in Drosophila. eLife 6. DOI


